wbia.algo.verif.torch package¶
Submodules¶
wbia.algo.verif.torch.fit_harness module¶
-
class
wbia.algo.verif.torch.fit_harness.FitHarness(model, train_loader, vali_loader=None, test_loader=None, criterion='cross_entropy', lr_scheduler='exp', optimizer_cls='Adam', class_weights=None, gpu_num=None, workdir=None)[source]¶ Bases:
object-
train_batch(input_batch)[source]¶ https://github.com/meetshah1995/pytorch-semseg/blob/master/train.py
-
wbia.algo.verif.torch.gpu_util module¶
wbia.algo.verif.torch.lr_schedule module¶
-
class
wbia.algo.verif.torch.lr_schedule.Exponential(init_lr=0.001, decay_rate=0.01, lr_decay_epoch=100)[source]¶ Bases:
objectDecay learning rate by a factor of decay_rate every lr_decay_epoch epochs.
Example
>>> # DISABLE_DOCTEST >>> from wbia.algo.verif.torch.lr_schedule import * >>> lr_scheduler = Exponential() >>> rates = np.array([lr_scheduler(i) for i in range(6)]) >>> target = np.array([1E-3, 1E-3, 1E-5, 1E-5, 1E-7, 1E-7]) >>> assert all(list(np.isclose(target, rates)))
wbia.algo.verif.torch.models module¶
wbia.algo.verif.torch.netmath module¶
-
class
wbia.algo.verif.torch.netmath.ContrastiveLoss(margin=1.0)[source]¶ Bases:
torch.nn.modules.module.ModuleContrastive loss function.
References
https://github.com/delijati/pytorch-siamese/blob/master/contrastive.py
- LaTeX:
- $(y E)^2 + ((1 - y) max(m - E, 0)^2)$
Example
>>> # DISABLE_DOCTEST >>> from wbia.algo.verif.siamese import * >>> vecs1, vecs2, label = testdata_siam_desc() >>> self = ContrastiveLoss() >>> ut.exec_func_src(self.forward, globals()) >>> func = self.forward >>> output = torch.nn.PairwiseDistance(p=2)(vecs1, vecs2) >>> loss2x, dist_l2 = ut.exec_func_src(self.forward, globals(), globals(), keys=['loss2x', 'dist_l2']) >>> ut.quit_if_noshow() >>> loss2x, dist_l2, label = map(np.array, [loss, dist_l2, label]) >>> label = label.astype(np.bool) >>> dist0_l2 = dist_l2[label] >>> dist1_l2 = dist_l2[~label] >>> loss0 = loss2x[label] / 2 >>> loss1 = loss2x[~label] / 2 >>> import wbia.plottool as pt >>> pt.plot2(dist0_l2, loss0, 'x', color=pt.TRUE_BLUE, label='imposter_loss', y_label='loss') >>> pt.plot2(dist1_l2, loss1, 'x', color=pt.FALSE_RED, label='genuine_loss', y_label='loss') >>> pt.gca().set_xlabel('l2-dist') >>> pt.legend() >>> ut.show_if_requested()
-
forward(output, label, weight=None)[source]¶ Defines the computation performed at every call.
Should be overridden by all subclasses.
Note
Although the recipe for forward pass needs to be defined within this function, one should call the
Moduleinstance afterwards instead of this since the former takes care of running the registered hooks while the latter silently ignores them.
-
class
wbia.algo.verif.torch.netmath.Criterions[source]¶ Bases:
wbia.algo.verif.torch.netmath.NetMathParamsA collection of standard and custom loss criterion
-
class
ContrastiveLoss(margin=1.0)¶ Bases:
torch.nn.modules.module.ModuleContrastive loss function.
References
https://github.com/delijati/pytorch-siamese/blob/master/contrastive.py
- LaTeX:
- $(y E)^2 + ((1 - y) max(m - E, 0)^2)$
Example
>>> # DISABLE_DOCTEST >>> from wbia.algo.verif.siamese import * >>> vecs1, vecs2, label = testdata_siam_desc() >>> self = ContrastiveLoss() >>> ut.exec_func_src(self.forward, globals()) >>> func = self.forward >>> output = torch.nn.PairwiseDistance(p=2)(vecs1, vecs2) >>> loss2x, dist_l2 = ut.exec_func_src(self.forward, globals(), globals(), keys=['loss2x', 'dist_l2']) >>> ut.quit_if_noshow() >>> loss2x, dist_l2, label = map(np.array, [loss, dist_l2, label]) >>> label = label.astype(np.bool) >>> dist0_l2 = dist_l2[label] >>> dist1_l2 = dist_l2[~label] >>> loss0 = loss2x[label] / 2 >>> loss1 = loss2x[~label] / 2 >>> import wbia.plottool as pt >>> pt.plot2(dist0_l2, loss0, 'x', color=pt.TRUE_BLUE, label='imposter_loss', y_label='loss') >>> pt.plot2(dist1_l2, loss1, 'x', color=pt.FALSE_RED, label='genuine_loss', y_label='loss') >>> pt.gca().set_xlabel('l2-dist') >>> pt.legend() >>> ut.show_if_requested()
-
forward(output, label, weight=None)¶ Defines the computation performed at every call.
Should be overridden by all subclasses.
Note
Although the recipe for forward pass needs to be defined within this function, one should call the
Moduleinstance afterwards instead of this since the former takes care of running the registered hooks while the latter silently ignores them.
-
class
-
class
wbia.algo.verif.torch.netmath.LRSchedules[source]¶ Bases:
wbia.algo.verif.torch.netmath.NetMathParamsA collection of standard and custom learning rate schedulers
-
class
wbia.algo.verif.torch.netmath.Optimizers[source]¶ Bases:
wbia.algo.verif.torch.netmath.NetMathParams-
class
Adam(params, lr=0.001, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)¶ Bases:
torch.optim.optimizer.OptimizerImplements Adam algorithm.
![\begin{aligned}
&\rule{110mm}{0.4pt} \\
&\textbf{input} : \gamma \text{ (lr)}, \beta_1, \beta_2
\text{ (betas)},\theta_0 \text{ (params)},f(\theta) \text{ (objective)} \\
&\hspace{13mm} \lambda \text{ (weight decay)}, \: amsgrad \\
&\textbf{initialize} : m_0 \leftarrow 0 \text{ ( first moment)},
v_0\leftarrow 0 \text{ (second moment)},\: \widehat{v_0}^{max}\leftarrow 0\\[-1.ex]
&\rule{110mm}{0.4pt} \\
&\textbf{for} \: t=1 \: \textbf{to} \: \ldots \: \textbf{do} \\
&\hspace{5mm}g_t \leftarrow \nabla_{\theta} f_t (\theta_{t-1}) \\
&\hspace{5mm}\textbf{if} \: \lambda \neq 0 \\
&\hspace{10mm} g_t \leftarrow g_t + \lambda \theta_{t-1} \\
&\hspace{5mm}m_t \leftarrow \beta_1 m_{t-1} + (1 - \beta_1) g_t \\
&\hspace{5mm}v_t \leftarrow \beta_2 v_{t-1} + (1-\beta_2) g^2_t \\
&\hspace{5mm}\widehat{m_t} \leftarrow m_t/\big(1-\beta_1^t \big) \\
&\hspace{5mm}\widehat{v_t} \leftarrow v_t/\big(1-\beta_2^t \big) \\
&\hspace{5mm}\textbf{if} \: amsgrad \\
&\hspace{10mm}\widehat{v_t}^{max} \leftarrow \mathrm{max}(\widehat{v_t}^{max},
\widehat{v_t}) \\
&\hspace{10mm}\theta_t \leftarrow \theta_{t-1} - \gamma \widehat{m_t}/
\big(\sqrt{\widehat{v_t}^{max}} + \epsilon \big) \\
&\hspace{5mm}\textbf{else} \\
&\hspace{10mm}\theta_t \leftarrow \theta_{t-1} - \gamma \widehat{m_t}/
\big(\sqrt{\widehat{v_t}} + \epsilon \big) \\
&\rule{110mm}{0.4pt} \\[-1.ex]
&\bf{return} \: \theta_t \\[-1.ex]
&\rule{110mm}{0.4pt} \\[-1.ex]
\end{aligned}](_images/math/e32d6b754636d7b367b785e24d78a81f4813fca4.png)
For further details regarding the algorithm we refer to Adam: A Method for Stochastic Optimization.
Parameters: - params (iterable) – iterable of parameters to optimize or dicts defining parameter groups
- lr (float, optional) – learning rate (default: 1e-3)
- betas (Tuple[float, float], optional) – coefficients used for computing running averages of gradient and its square (default: (0.9, 0.999))
- eps (float, optional) – term added to the denominator to improve numerical stability (default: 1e-8)
- weight_decay (float, optional) – weight decay (L2 penalty) (default: 0)
- amsgrad (boolean, optional) – whether to use the AMSGrad variant of this algorithm from the paper On the Convergence of Adam and Beyond (default: False)
-
step(closure=None)¶ Performs a single optimization step.
Parameters: closure (callable, optional) – A closure that reevaluates the model and returns the loss.
-
class
SGD(params, lr=<required parameter>, momentum=0, dampening=0, weight_decay=0, nesterov=False)¶ Bases:
torch.optim.optimizer.OptimizerImplements stochastic gradient descent (optionally with momentum).
![\begin{aligned}
&\rule{110mm}{0.4pt} \\
&\textbf{input} : \gamma \text{ (lr)}, \: \theta_0 \text{ (params)}, \: f(\theta)
\text{ (objective)}, \: \lambda \text{ (weight decay)}, \\
&\hspace{13mm} \:\mu \text{ (momentum)}, \:\tau \text{ (dampening)},\:nesterov\\[-1.ex]
&\rule{110mm}{0.4pt} \\
&\textbf{for} \: t=1 \: \textbf{to} \: \ldots \: \textbf{do} \\
&\hspace{5mm}g_t \leftarrow \nabla_{\theta} f_t (\theta_{t-1}) \\
&\hspace{5mm}\textbf{if} \: \lambda \neq 0 \\
&\hspace{10mm} g_t \leftarrow g_t + \lambda \theta_{t-1} \\
&\hspace{5mm}\textbf{if} \: \mu \neq 0 \\
&\hspace{10mm}\textbf{if} \: t > 1 \\
&\hspace{15mm} \textbf{b}_t \leftarrow \mu \textbf{b}_{t-1} + (1-\tau) g_t \\
&\hspace{10mm}\textbf{else} \\
&\hspace{15mm} \textbf{b}_t \leftarrow g_t \\
&\hspace{10mm}\textbf{if} \: nesterov \\
&\hspace{15mm} g_t \leftarrow g_{t-1} + \mu \textbf{b}_t \\
&\hspace{10mm}\textbf{else} \\[-1.ex]
&\hspace{15mm} g_t \leftarrow \textbf{b}_t \\
&\hspace{5mm}\theta_t \leftarrow \theta_{t-1} - \gamma g_t \\[-1.ex]
&\rule{110mm}{0.4pt} \\[-1.ex]
&\bf{return} \: \theta_t \\[-1.ex]
&\rule{110mm}{0.4pt} \\[-1.ex]
\end{aligned}](_images/math/88be97cfbb089937c5484c4965ac38c73c294173.png)
Nesterov momentum is based on the formula from On the importance of initialization and momentum in deep learning.
Parameters: - params (iterable) – iterable of parameters to optimize or dicts defining parameter groups
- lr (float) – learning rate
- momentum (float, optional) – momentum factor (default: 0)
- weight_decay (float, optional) – weight decay (L2 penalty) (default: 0)
- dampening (float, optional) – dampening for momentum (default: 0)
- nesterov (bool, optional) – enables Nesterov momentum (default: False)
Example
>>> optimizer = torch.optim.SGD(model.parameters(), lr=0.1, momentum=0.9) >>> optimizer.zero_grad() >>> loss_fn(model(input), target).backward() >>> optimizer.step()
Note
The implementation of SGD with Momentum/Nesterov subtly differs from Sutskever et. al. and implementations in some other frameworks.
Considering the specific case of Momentum, the update can be written as
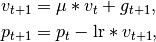
where
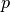 ,
, 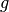 ,
,  and
and 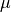 denote the
parameters, gradient, velocity, and momentum respectively.
denote the
parameters, gradient, velocity, and momentum respectively.This is in contrast to Sutskever et. al. and other frameworks which employ an update of the form
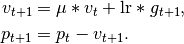
The Nesterov version is analogously modified.
-
step(closure=None)¶ Performs a single optimization step.
Parameters: closure (callable, optional) – A closure that reevaluates the model and returns the loss.
-
class
wbia.algo.verif.torch.old_harness module¶
wbia.algo.verif.torch.siamese module¶
wbia.algo.verif.torch.train_main module¶
-
class
wbia.algo.verif.torch.train_main.LabeledPairDataset(img1_fpaths, img2_fpaths, labels, transform='default')[source]¶ Bases:
torch.utils.data.dataset.Dataset- transform=transforms.Compose([
- transforms.Scale(224), transforms.ToTensor(), torchvision.transforms.Normalize([0.5, 0.5, 0.5], [0.225, 0.225, 0.225])
]
- Ignore:
>>> from wbia.algo.verif.torch.train_main import * >>> from wbia.algo.verif.vsone import * # NOQA >>> pblm = OneVsOneProblem.from_empty('PZ_MTEST') >>> ibs = pblm.infr.ibs >>> pblm.load_samples() >>> samples = pblm.samples >>> samples.print_info() >>> xval_kw = pblm.xval_kw.asdict() >>> skf_list = pblm.samples.stratified_kfold_indices(**xval_kw) >>> train_idx, test_idx = skf_list[0] >>> aids1, aids2 = pblm.samples.aid_pairs[train_idx].T >>> labels = pblm.samples['match_state'].y_enc[train_idx] >>> labels = (labels == 1).astype(np.int64) >>> chip_config = {'resize_dim': 'wh', 'dim_size': (224, 224)} >>> img1_fpaths = ibs.depc_annot.get('chips', aids1, read_extern=False, colnames='img', config=chip_config) >>> img2_fpaths = ibs.depc_annot.get('chips', aids2, read_extern=False, colnames='img', config=chip_config) >>> self = LabeledPairDataset(img1_fpaths, img2_fpaths, labels) >>> img1, img2, label = self[0]